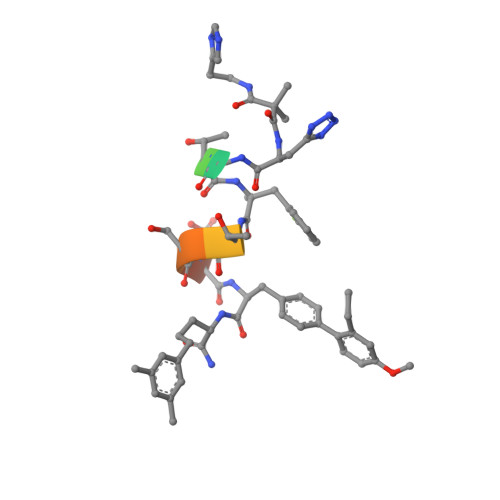
Crystal structure of the GLP-1 receptor bound to a peptide agonist.
Jazayeri, A., Rappas, M., Brown, A.J.H., Kean, J., Errey, J.C., Robertson, N.J., Fiez-Vandal, C., Andrews, S.P., Congreve, M., Bortolato, A., Mason, J.S., Baig, A.H., Teobald, I., Dore, A.S., Weir, M., Cooke, R.M., Marshall, F.H.(2017) Nature 546: 254-258
- PubMed: 28562585 Search on PubMed
- DOI: https://doi.org/10.1038/nature22800
- Primary Citation Related Structures:
5NX2 - PubMed Abstract:
Glucagon-like peptide 1 (GLP-1) regulates glucose homeostasis through the control of insulin release from the pancreas. GLP-1 peptide agonists are efficacious drugs for the treatment of diabetes. To gain insight into the molecular mechanism of action of GLP-1 peptides, here we report the crystal structure of the full-length GLP-1 receptor bound to a truncated peptide agonist. The peptide agonist retains an α-helical conformation as it sits deep within the receptor-binding pocket. The arrangement of the transmembrane helices reveals hallmarks of an active conformation similar to that observed in class A receptors. Guided by this structural information, we design peptide agonists with potent in vivo activity in a mouse model of diabetes.
- Heptares Therapeutics Ltd, BioPark, Broadwater Road, Welwyn Garden City, Hertfordshire AL7 3AX, UK.
Organizational Affiliation: